Speaker
Description
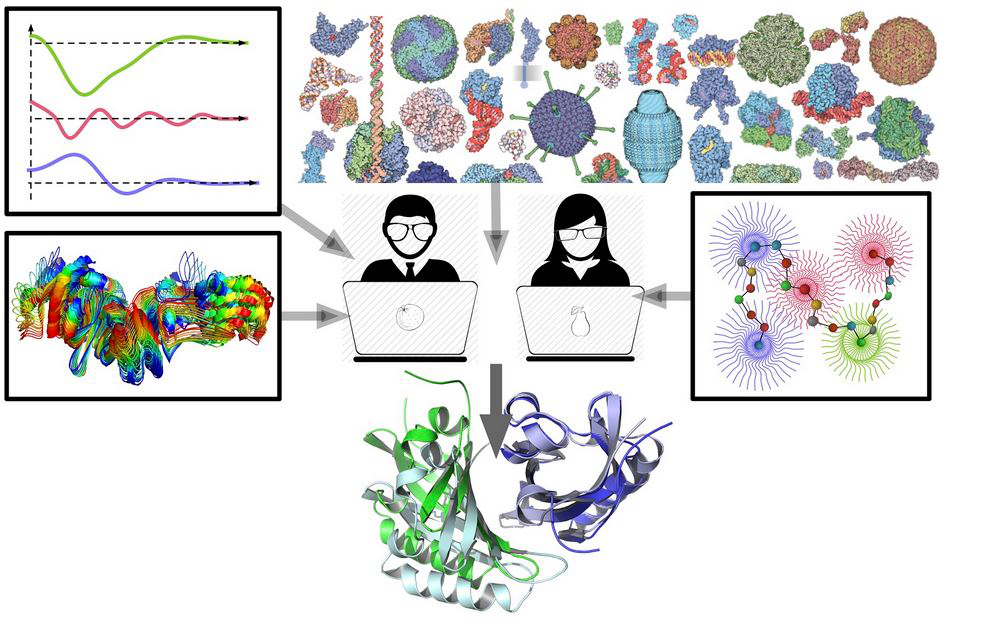
Given a molecular structure, it can be represented as a set of atomic balls, each ball having a van der Waals radius corresponding to the atom type. A ball can be assigned a region of space that contains all the points that are closer (or equally close) to that ball than to any other. Such a region is called a Voronoi cell and the partitioning of space into Voronoi cells is called Voronoi tessellation or Voronoi diagram. Two adjacent Voronoi cells share a set of points that form a surface called a Voronoi face. A Voronoi face can be viewed as a geometric representation of a contact between two atoms. The Voronoi cells of atomic balls may be constrained inside the boundaries defined by the solvent accessible surface of the same balls. The constrained Voronoi cells and their faces are remarkably versatile structural descriptors of atoms and their interactions. Another example of tessellation-derived features are empty tangent spheres corresponding to the vertices of the Voronoi cells. This talk will be focused on some of the protein structural analysis and assessment algorithms that are built upon the aforementioned Voronoi tessellation-derived descriptors.