Speaker
Description
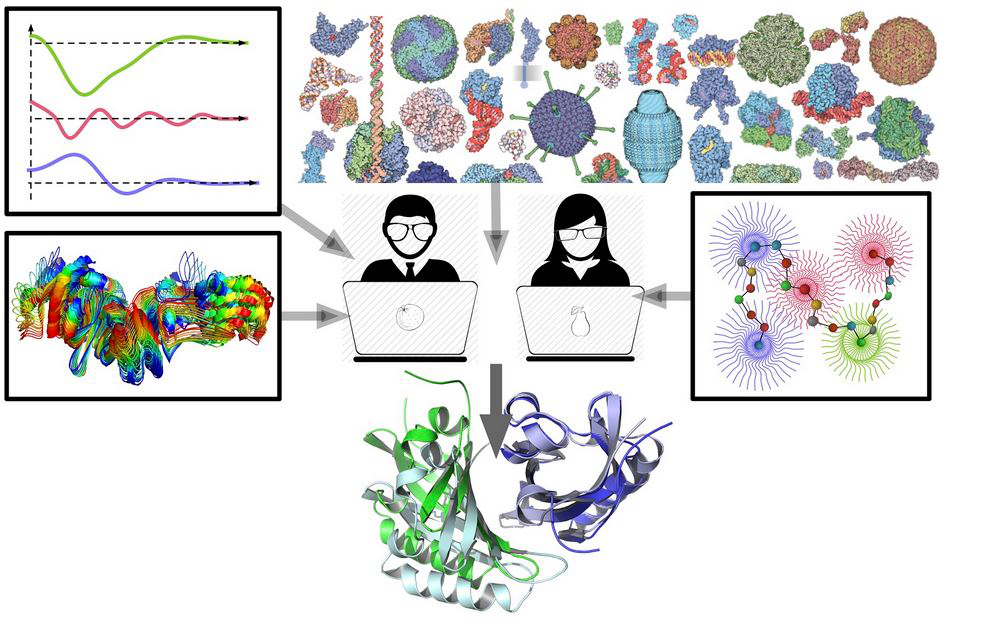
Understanding how proteins and complexes move in solution remains a major challenge in structural biology. Biological macromolecules are machines that adopt a variety of structural conformations and efficiently exploring this structural space in a comprehensive and meaningful way can not be achieved solely using the solid-state methods cryo-EM and macromolecular crystallography (MX). Small X-ray scattering (SAXS) measurements of biological macromolecular particles in solution provides a resolution-limited, structural assessment of the thermodynamic ensemble. In a SAXS measurement, all conformations will be sampled albeit with a severe loss resolution and correspondence. Nonetheless,, SAXS observations made at higher resolutions imply a greater detail in the structural measurement. Here, I will present an approach to understanding SAXS data using Information Theory (IT). I will show that the Information theory framework provides a pathway for exploring conformational space when integrated with computational methods such as molecular dynamics. The IT framework can be used to develop structural modeling algorithms for shape determination and docking and I will show the IT approach can be used in antibody-antigen studies to uniquely determine the structure of the complex in the solution state. The IT approach suggests a general method for refining atomistic models derived from homology models or prediction.